When size matters: selection of training sets for support vector machines
- Processes, standards and quality
- Technologies
- Others
The era of big data is here and now. How to efficiently train support vector machines from massively large real-life datasets?
The Background
The amount of data produced every day grows tremendously in most real-life domains, including medical imaging, genomics, text categorisation, computational biology, and many others. Although it appears beneficial at the first glance (more data could mean more possibilities of extracting and revealing useful underlying knowledge), handling massively large datasets became a challenging issue and attracts research attention, especially in the era of big data. This big data revolution affected many research fields, including statistics, machine learning, parallel computing, and computer systems in general [1].
Storing and analysing the acquired historical information should allow predicting the label of an incoming (unseen) feature vector, containing some quantified features of a given data example. If the labels are categorical, then we are to tackle the classification task (it’s regression otherwise). In this note, we will consider the binary (two-class) classification. There are a plethora of various classifiers available at hand, ranging from basic nearest-neighbours based systems to neural networks, non-linear kernel machines (you do remember the kernel trick [2], don’t you?), and deep learning methods, which are super-hot buzzwords at the moment.
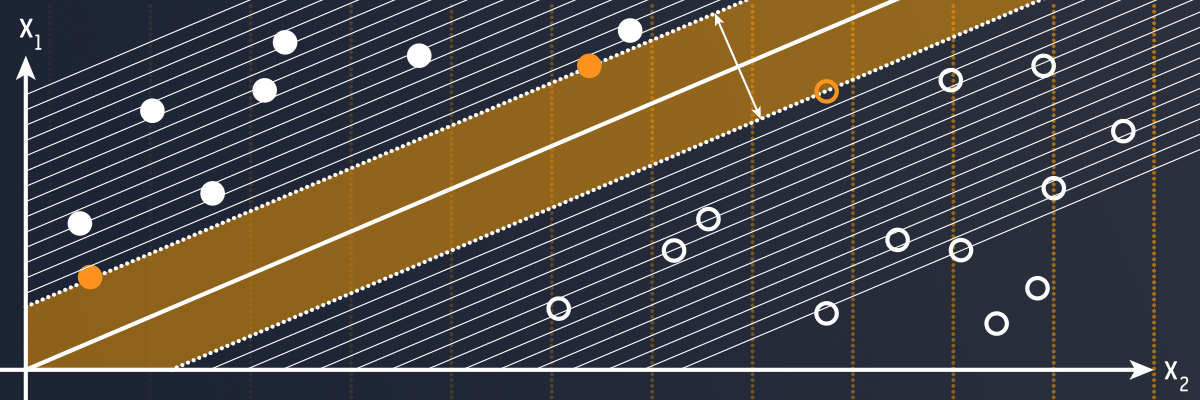
Support vector machine (SVM) is a supervised classifier which has been applied for solving a wide range of pattern recognition problems. However, training of SVMs may easily become their bottleneck, because of its time and memory requirements (O(t3) and O(t2), respectively, where t denotes the cardinality of the training set). Enduring this issue is pivotal to make SVMs applicable in real-life scenarios. Can we deal with that efficiently?
What Can We Do?
Training of SVMs involves searching for a subset of the entire training set T encompassing vectors which define the position of the decision hyperplane (these vectors are called support vectors, SVs). Without diving into details, it’s a constrained quadratic programming optimisation problem. Two main streams of research towards dealing with this time-consuming SVM training can be distinguished. The first stream includes algorithms which aim at boosting the training process itself, whereas the other – at retrieving significantly smaller refined T’s beforehand. Since only a small subset of T is selected as SVs, training SVMs with subsets of T containing only SVs should give the same model as if they were trained with T. Why do not exploit this observation as best as possible?
Although the first group of methods is being quite intensively developed (look at reduced SVMs, parallel methods, or online twin independent SVMs, the last being hot off the press [3]), they often suffer from large memory requirements. The second group of approaches can be further divided into several subgroups: statistical methods, clustering-based techniques, not-so-trivial random sampling algorithms, and those looking at geometrical characteristics of T in search of its valuable subsets. What was our remedy?
The Remedy: Evolutionary Algorithms
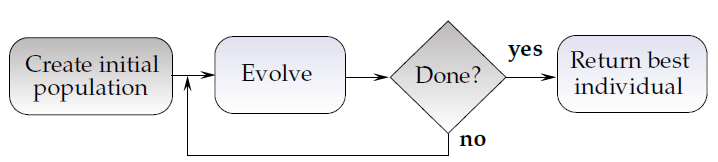
Retrieving refined SVM training sets using evolutionary algorithms (EAs) was our research subject for the last years. Our efforts were carried out at the Silesian University of Technology within a research project, which Michał Kawulok and myself were granted from the Polish Ministry of Science and Higher Education. Starting from relatively simple genetic algorithms (GAs) [4] (this approach was presented at S+SSPR 2012 in Hiroshima, Japan), through two adaptive GAs [5] [6] (these algorithms were presented at EVOStar 2014 in Granada, Spain, and at IBERAMIA 2014 in Santiago, Chile, respectively), towards memetic approaches [7] (presented at GECCO 2014 in Vancouver, Canada), and adaptive memetic algorithms [8] (the hybrids of GAs and refinement procedures exploiting all available knowledge to improve individuals during the evolution), we showed that they outperform the state of the art. The idea is simple: create a random population of refined training sets (individuals) sampled from T and evolve them (as shown in Figure 1). Are we done yet? Absolutely not – that’s the beginning.
There are a bunch of questions yet to be answered: what is the best selection scheme, crossover operator, mutation rate, or – most importantly – the size of individuals (chromosomes). This size is independent from the cardinality of T. All of the aforementioned parameters can (and in fact do) affect the search progress and convergence capabilities of EAs. Evolutionary techniques are inherently dynamic, thus the optimal value of a given parameter will most likely vary in time. This is where the adaptation schemes come to play: monitoring of the temporal changes of the population characteristics (e.g., the fitness of the best individual or the population diversity) is quite handy and it may trigger the parameter updates. The exploration-exploitation trade-off looks certainly more manageable then.
Wasting the acquired knowledge is irresponsible and this applies to algorithms as well. Some individuals during the evolution are well-fitted, some are poorly-fitted – that’s usual. This is, however, quite an important bit of information as long as we are able to look at those chromosomes in more detail. Yes, we are.
Determining the fitness of an individual consists in training an SVM using the corresponding refined set and finding the area under the receiver operating characteristic curve obtained for T. Hence, we know which vectors from T are important (they are selected as SVs), and are likely to be selected as SVs again. These vectors are then used to educate other individuals (the weak vectors are replaced with these potentially valuable ones with some probability). It is never too late to learn.
Can the evolution be enhanced even before it starts? Yes, it can. On the one hand, creating an initial population, which is not diverse, can easily jeopardise the optimisation process. On the other hand, if this population is well-established and the individuals are of high quality from the very beginning, then the evolution may converge to the desired refined sets faster. This was indeed the case in our adaptive memetic algorithm, in which the principal component analysis based pre-processing technique was applied to extract a subset of T before the optimisation – these vectors were utilised to not only create the initial population, but also to introduce new genetic material during the evolution.
(Not Only) The Size Does Matter
It’s really easy to imagine training sets with billions of feature vectors (and the number of features can be huge, too). As already shown, this size does matter indeed when a large T is to be fed into the SVM training procedure. What is important, lowering the cardinality of T affects the classification time as well (ideally, without deteriorating the classification score) – this time linearly depends on the number of SVs. In practice, the smaller training sets induce the smaller numbers of SVs. This is how to kill two birds with just one stone.
The size is not the only issue to tackle. Training set vectors may be mislabelled (or weakly labelled) – this situation is visualised in Figure 2 (black and white pixels represent vectors from T belonging to two classes, whereas mislabelled vectors are rendered in red). It can be a significant problem while extracting appropriate refined SVM training sets and should be addressed separately (e.g., by the label evolution, as we proposed in [9]).
SVMs are a very powerful tool, but should be used with care. Understanding the underlying training process of SVMs (and other classifiers too) is essential. That should be the very first step of applying a given classification engine to certain pattern recognition tasks. This knowledge helps address potential shortcomings and pitfalls of classifiers efficiently (without unnecessary wresting and tweaking). The life is much easier then.
Epilogue
I will present our freshest adaptive memetic algorithm (published in the Neurocomputing journal) at the GECCO 2016 Hot-off-the-Press session in Denver, USA (July, 20-24th, 2016). Shall we meet there?
The results of this work will be further exploited in the ECONIB project (extracting biomarkers from dynamic medical images). For more details, see https://better.future-processing.com/. Will you join us?
Acknowledgements
This work was supported by the Polish Ministry of Science and Higher Education under research grant no. IP2012 026372 from the Science Budget 2013 – 2015. The research was performed on the infrastructure supported by POIG.02.03.01-24-099/13 grant: “GeCONiI – Upper Silesian Centre for Computational Science and Engineering”.
Bibliography
[1] S. Haykin, S. Wright and Y. Bengio, „Big Data: Theoretical Aspects,” Proceedings of the IEEE, pp. 8-10, 2016.
[2] C. Cortes and V. Vapnik, „Support-vector networks,” Machine Learning, vol. 20, no. 3, pp. 273-297, 1995.
[3] F. Alamdar, S. Ghane and A. Amiri, „On-line twin independent support vector machines,” Neurocomputing, vol. 186, pp. 8-21, 2016.
[4] M. Kawulok and J. Nalepa, „Support vector machines training data selection using a genetic algorithm,” in Structural, Syntactic, and Statistical Pattern Recognition, Springer Berlin Heidelberg, 2012, pp. 557-565.
[5] J. Nalepa and M. Kawulok, “Adaptive genetic algorithm to select training data for support vector machines,” in Applications of Evolutionary Computation, Springer Berlin Heidelberg, 2014, pp. 514-525.
[6] M. Kawulok and J. Nalepa, “Dynamically Adaptive Genetic Algorithm to Select Training Data for SVMs,” in Advances in Artificial Intelligence – IBERAMIA 2014, Springer International Publishing, 2014, pp. 242-254.
[7] J. Nalepa and M. Kawulok, “A memetic algorithm to select training data for support vector machines,” in Proceedings of the 2014 conference on Genetic and evolutionary computation (GECCO 2014), Vancouver, Canada, 2014.
[8] J. Nalepa and M. Kawulok, “Adaptive memetic algorithm enhanced with data geometry analysis to select training data for SVMs,” Neurocomputing, vol. 185, p. 113–132, 2016.
[9] M. Kawulok and J. Nalepa, “Towards Robust SVM Training from Weakly Labeled Large Data Sets,” in Asian Conference on Pattern Recognition (ACPR 2015), Kuala Lumpur, 2015.
![Figure 2 - Examples of mislabelled vectors in T. This figure comes from our paper [9] - Future Processing](https://kariera.future-processing.pl//wp-content/uploads/2016/04/Figure-2-Examples-of-mislabelled-vectors-in-T.-This-figure-comes-from-our-paper-9-Future-Processing.png)